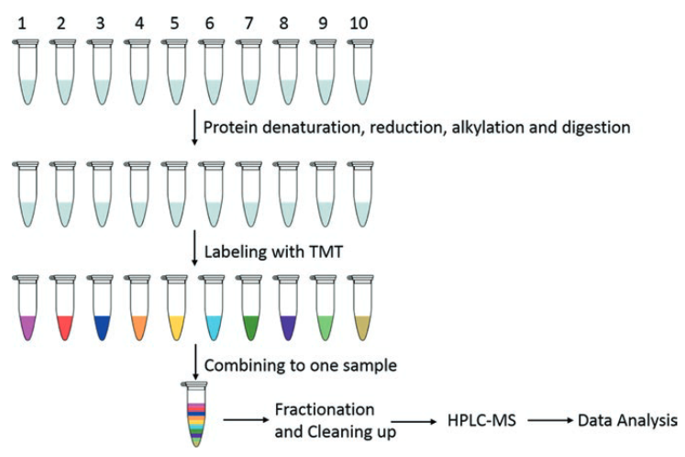
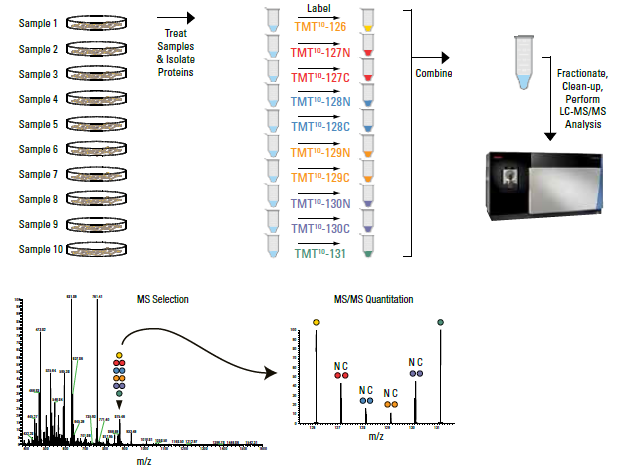
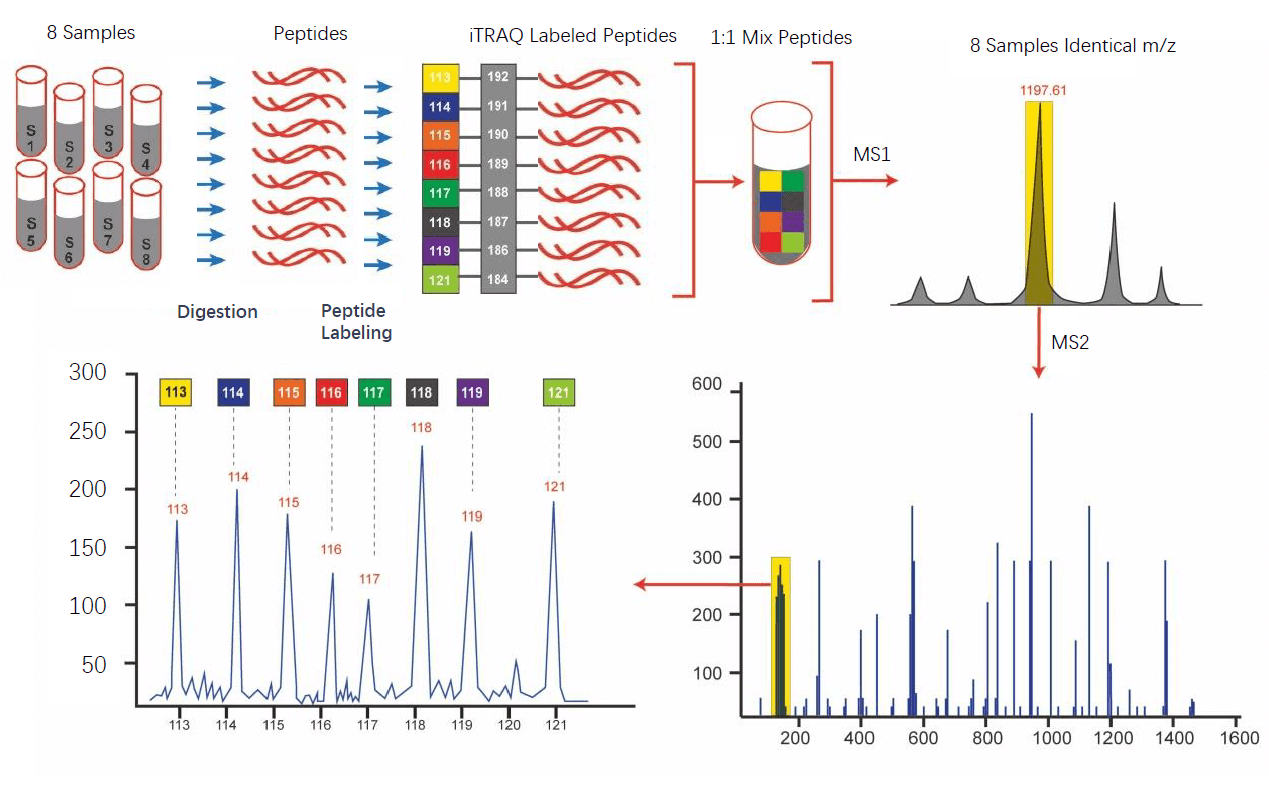
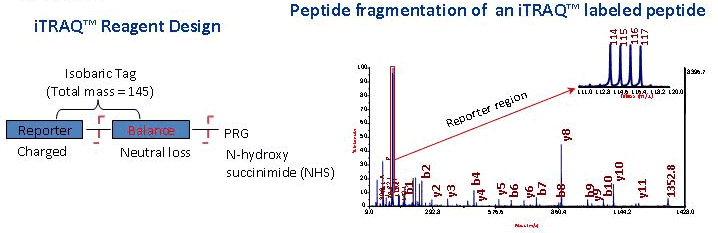
An efficient solution for resolving iTRAQ and TMT channel cross‐talk - Searle - 2020 - Journal of Mass Spectrometry - Wiley Online Library

Label-free and isobaric tandem mass tag (TMT) multiplexed quantitative proteomic data of two contrasting rice cultivars exposed to drought stress and recovery - ScienceDirect
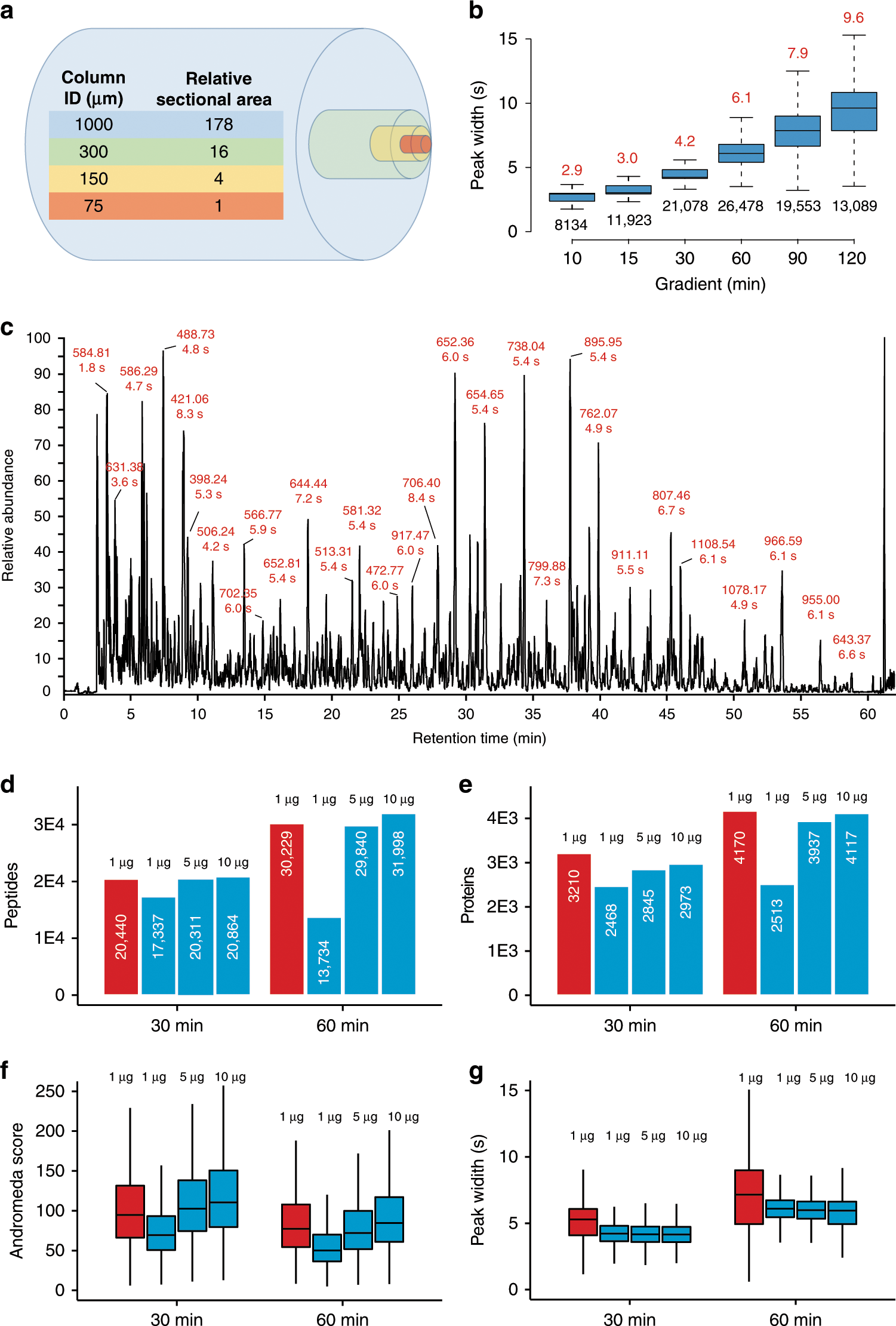
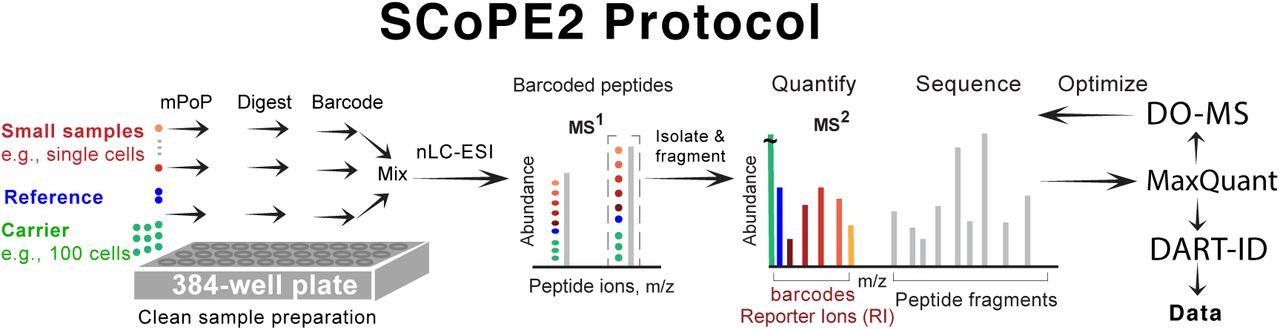
Robust, reproducible and quantitative analysis of thousands of proteomes by micro-flow LC–MS/MS | Nature Communications

Figure 1 from Proteome-Wide Evaluation of Two Common Protein Quantification Methods. | Semantic Scholar

Improving Quantitative Accuracy and Precision of Isobaric Labeling Strategies for Quantitative Proteomics Using Multistage (MS3) Mass Spectrometry | American Laboratory
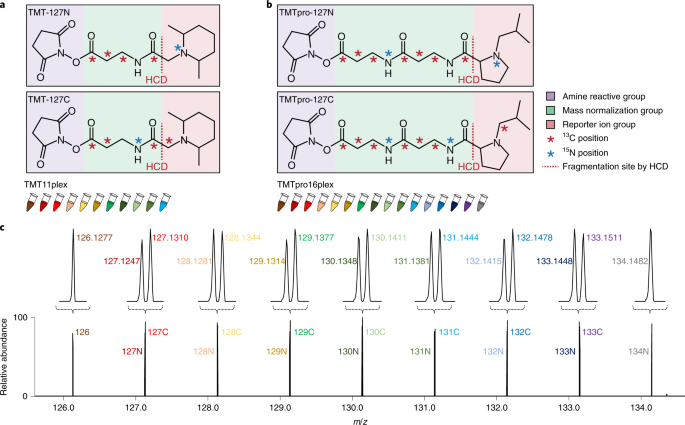
TMTpro reagents: a set of isobaric labeling mass tags enables simultaneous proteome-wide measurements across 16 samples | Nature Methods
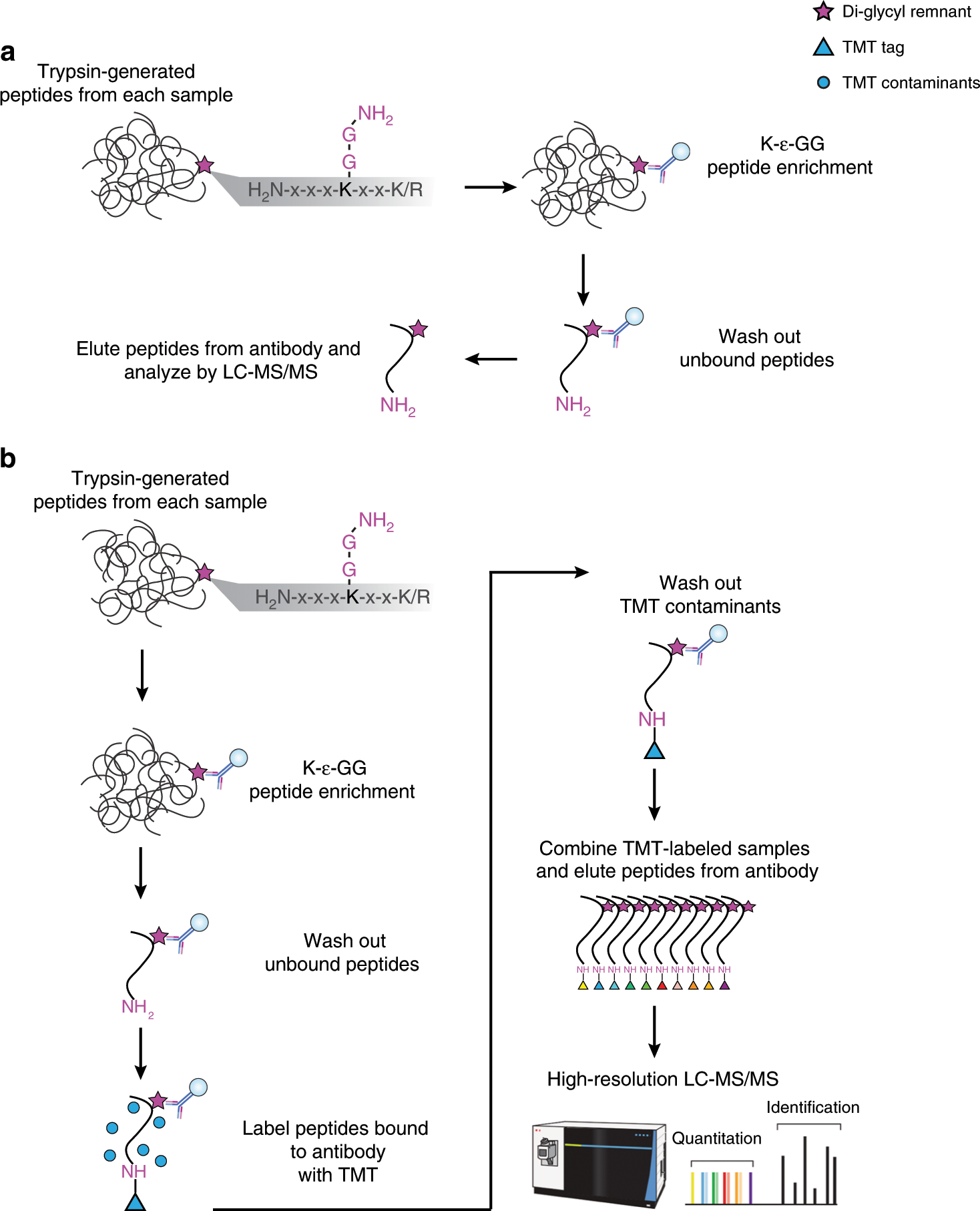
Rapid and deep-scale ubiquitylation profiling for biology and translational research | Nature Communications
![TMT Labeling for the Masses: A Robust and Cost-efficient, In-solution Labeling Approach*[S] - Molecular & Cellular Proteomics TMT Labeling for the Masses: A Robust and Cost-efficient, In-solution Labeling Approach*[S] - Molecular & Cellular Proteomics](https://els-jbs-prod-cdn.jbs.elsevierhealth.com/cms/attachment/a3452a3d-b954-4b77-b271-1d888f43c76c/gr3_lrg.jpg)
TMT Labeling for the Masses: A Robust and Cost-efficient, In-solution Labeling Approach*[S] - Molecular & Cellular Proteomics

Streamlined Tandem Mass Tag (SL-TMT) Protocol: An Efficient Strategy for Quantitative (Phospho)proteome Profiling Using Tandem Mass Tag-Synchronous Precursor Selection-MS3. - Abstract - Europe PMC