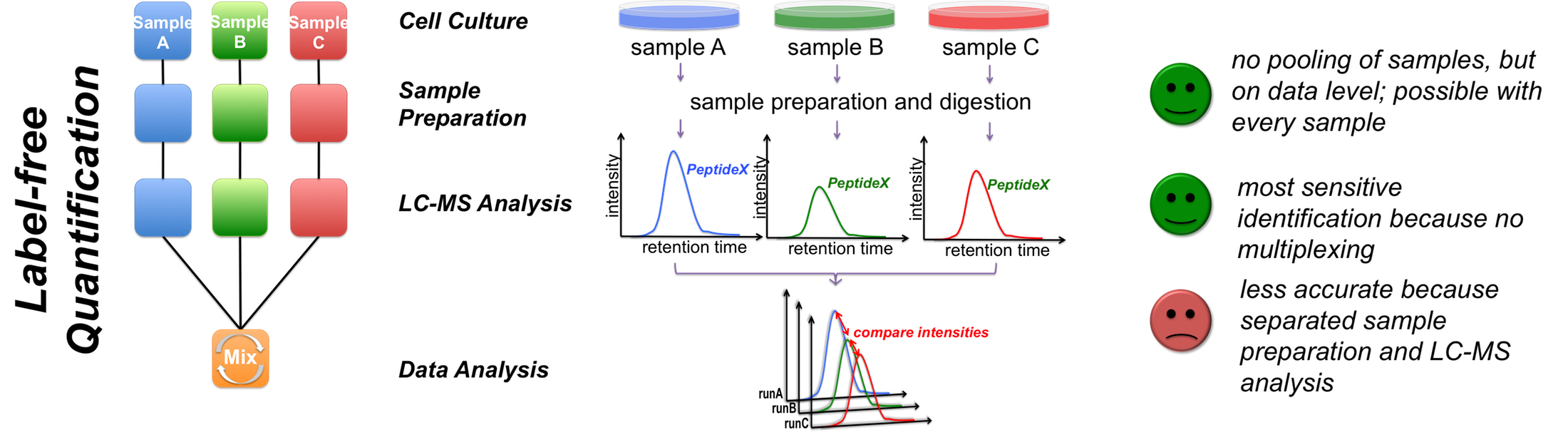
Accurate Proteome-wide Label-free Quantification by Delayed Normalization and Maximal Peptide Ratio Extraction, Termed MaxLFQ* - Molecular & Cellular Proteomics
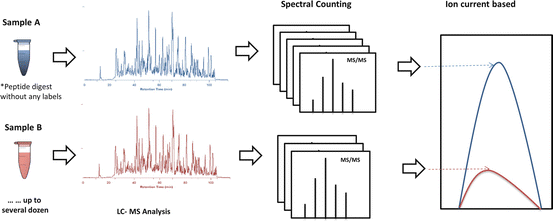
Labeling and Label-Free Shotgun Proteomics Quantification in the Research of Cardiovascular Diseases | SpringerLink
PLOS ONE: Comparative Analysis of Label-Free and 8-Plex iTRAQ Approach for Quantitative Tissue Proteomic Analysis

IonStar enables high-precision, low-missing-data proteomics quantification in large biological cohorts | PNAS

Mass spectrometry–based measurements. (a) Sample processing. Label-free... | Download Scientific Diagram

Robust Summarization and Inference in Proteome-wide Label-free Quantification - Molecular & Cellular Proteomics

Label-free quantitative proteomics of the lysine acetylome in mitochondria identifies substrates of SIRT3 in metabolic pathways | PNAS

A Review on Quantitative Multiplexed Proteomics - Pappireddi - 2019 - ChemBioChem - Wiley Online Library

Differences between label free quantification (A) and quantification by... | Download Scientific Diagram

Quantitative Mass Spectrometry-Based Approaches in Cardiovascular Research | Circulation: Cardiovascular Genetics
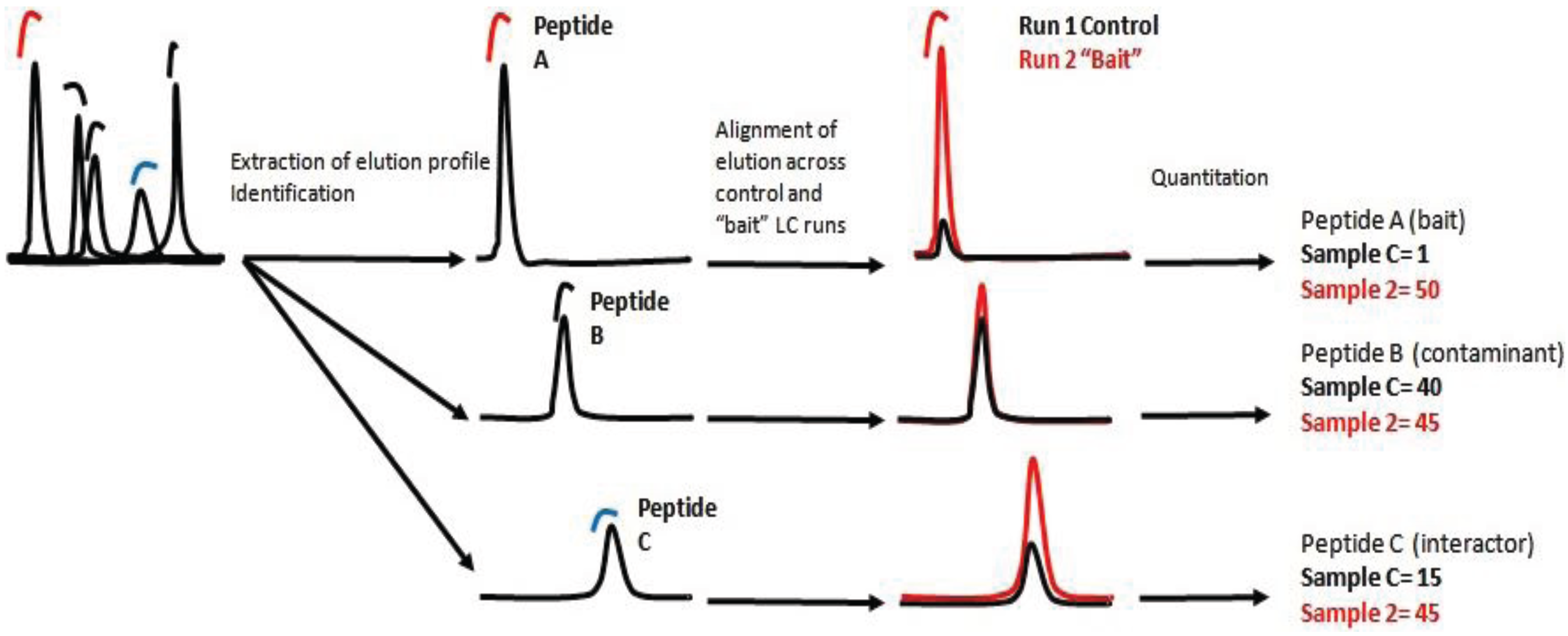
Biology | Free Full-Text | On-Beads Digestion in Conjunction with Data-Dependent Mass Spectrometry: A Shortcut to Quantitative and Dynamic Interaction Proteomics | HTML
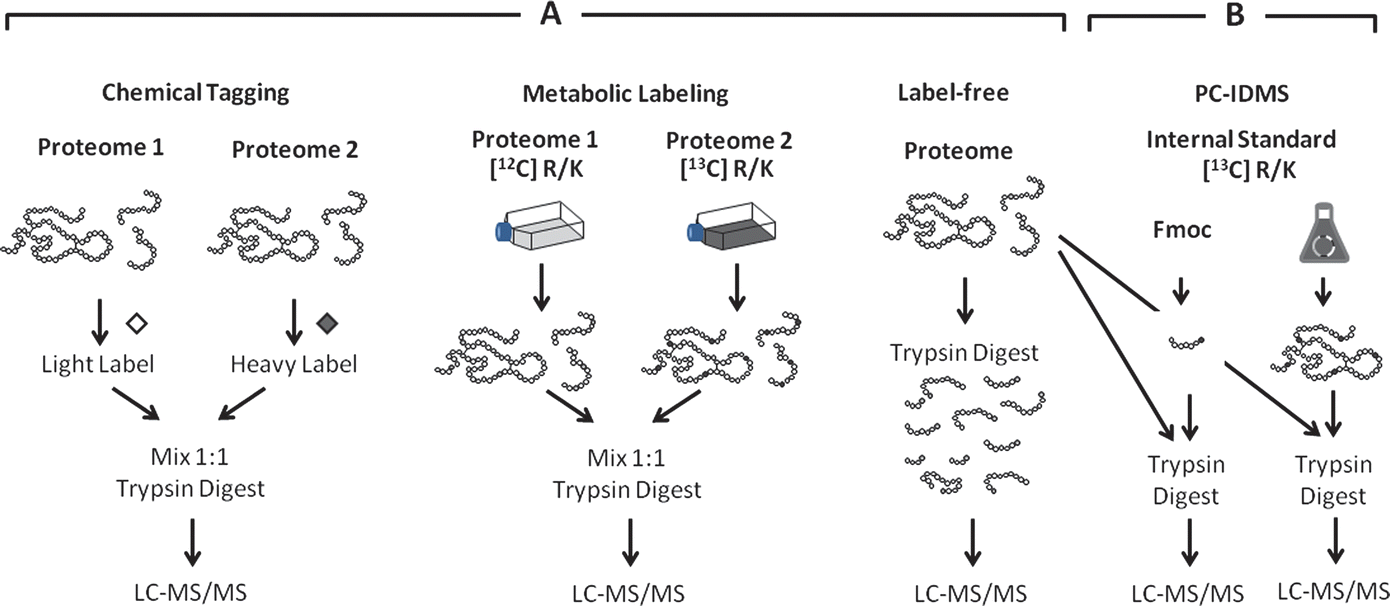

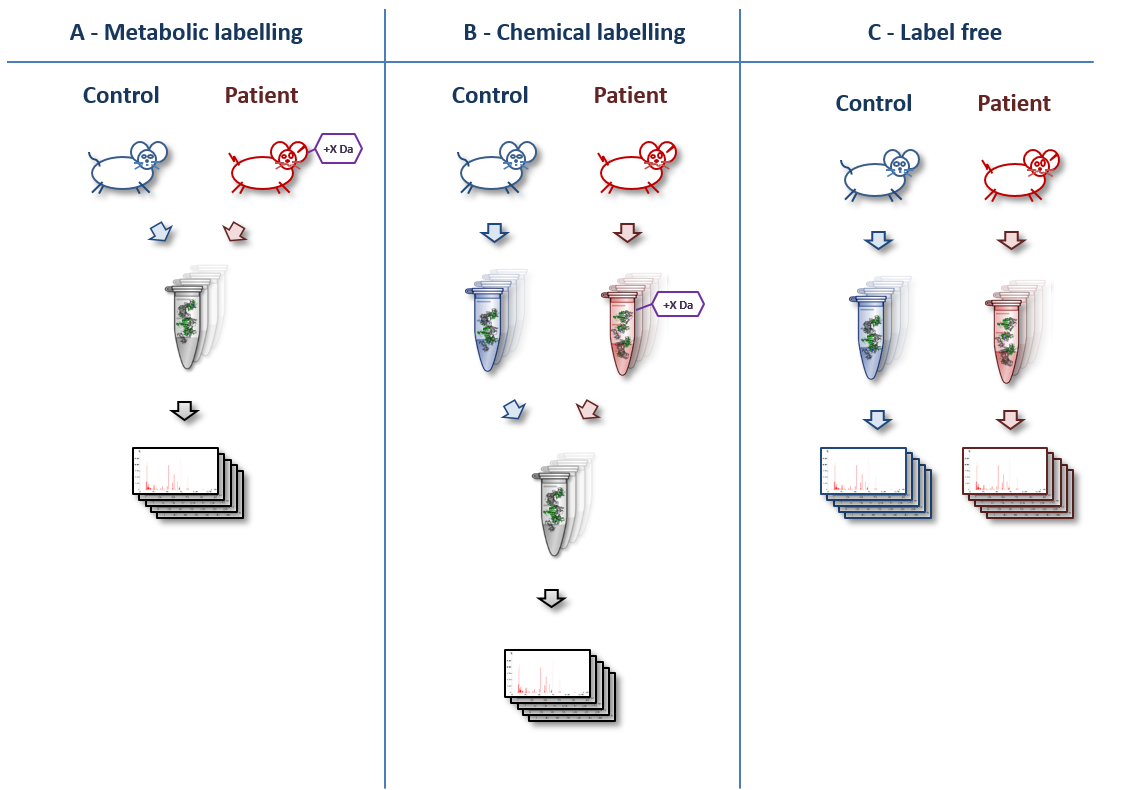
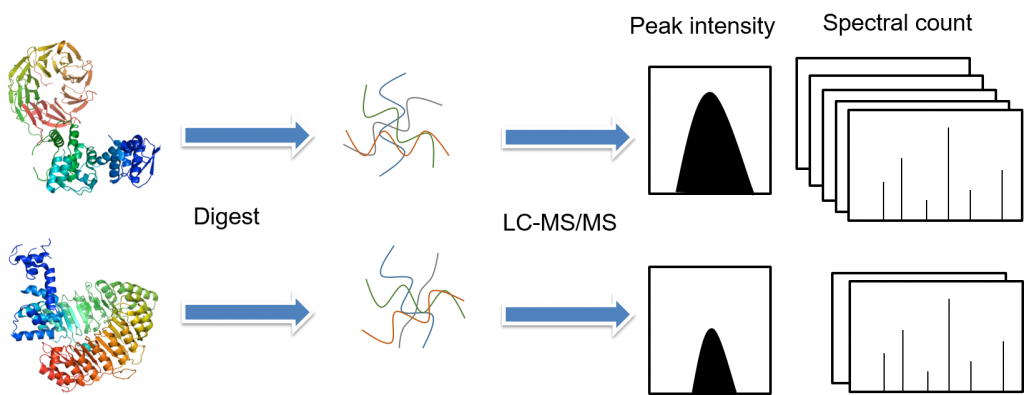



.jpg)