Mutational analysis of the neurexin/neuroligin complex reveals essential and regulatory components | PNAS
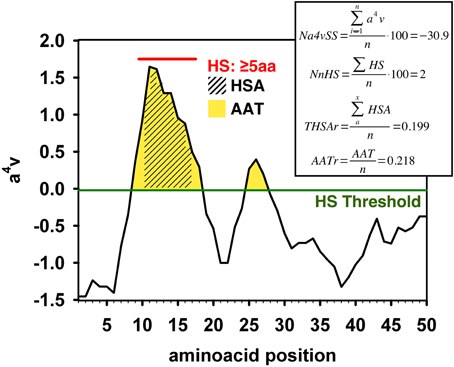
AGGRESCAN: a server for the prediction and evaluation of "hot spots" of aggregation in polypeptides | BMC Bioinformatics | Full Text
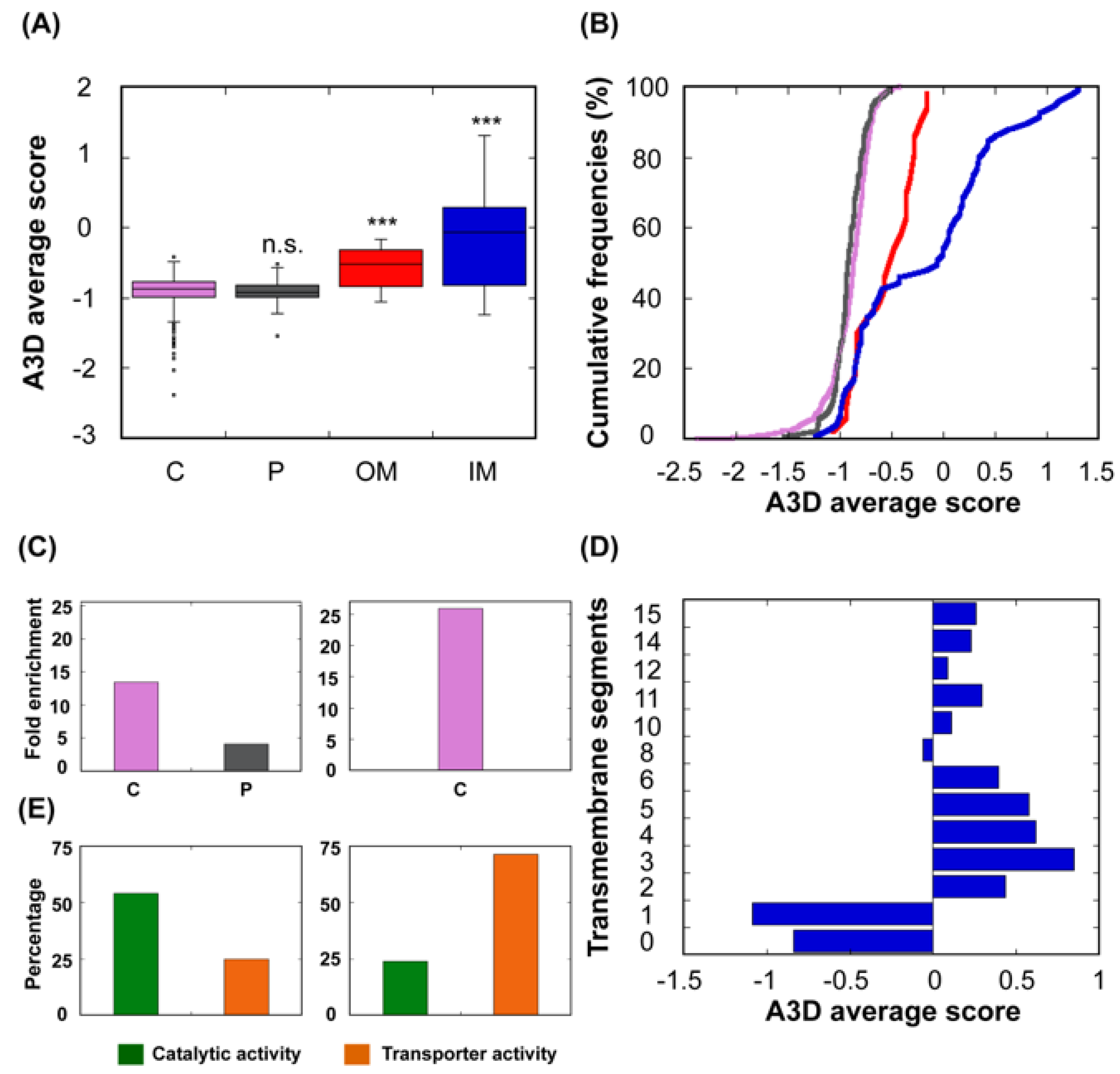
Cells | Free Full-Text | Computational Assessment of Bacterial Protein Structures Indicates a Selection Against Aggregation | HTML

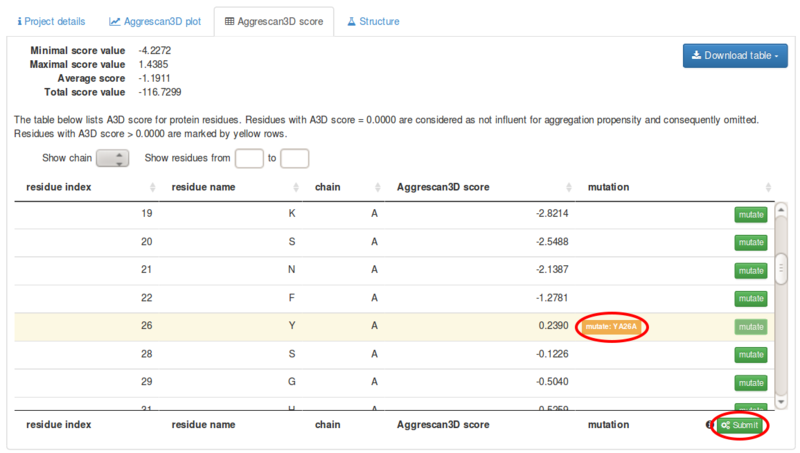
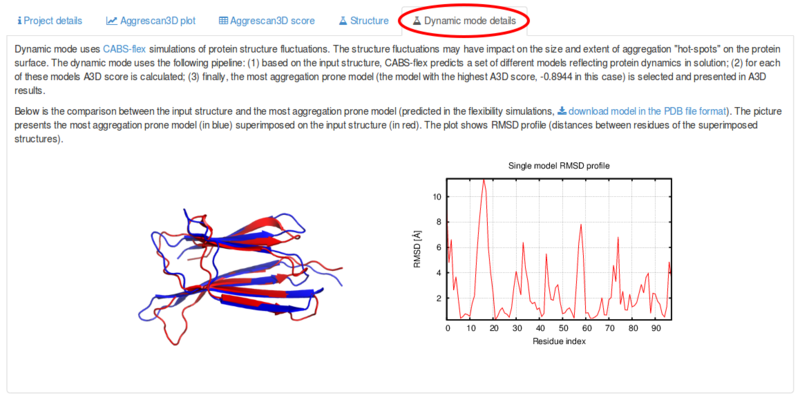
Figure 2 from Aggrescan3D (A3D) 2.0: prediction and engineering of protein solubility | Semantic Scholar
![PDF] AGGRESCAN3D (A3D): server for prediction of aggregation properties of protein structures | Semantic Scholar PDF] AGGRESCAN3D (A3D): server for prediction of aggregation properties of protein structures | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/bc8d0acd077eed82b977fbf9d706415235ef1fe6/7-Figure4-1.png)
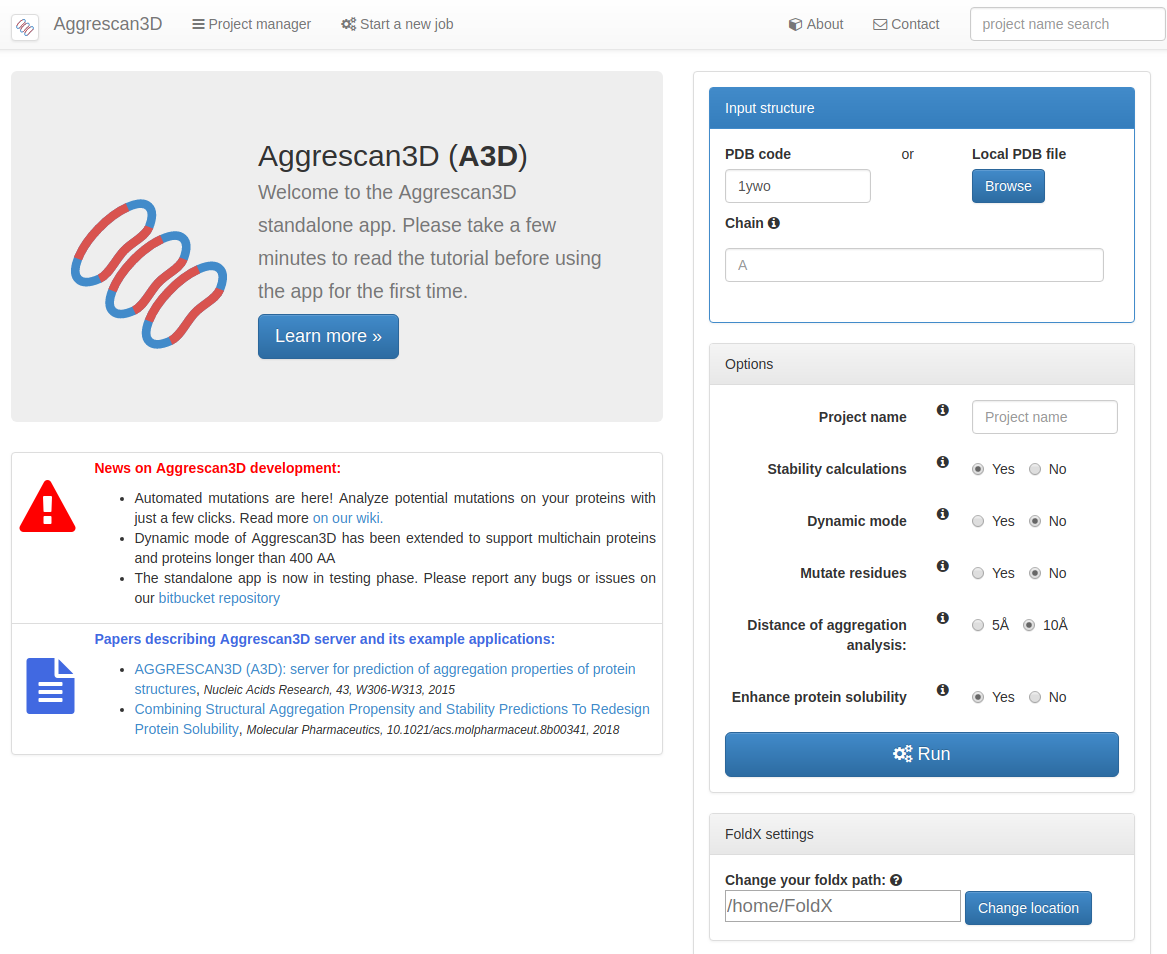
PDF] AGGRESCAN3D (A3D): server for prediction of aggregation properties of protein structures | Semantic Scholar

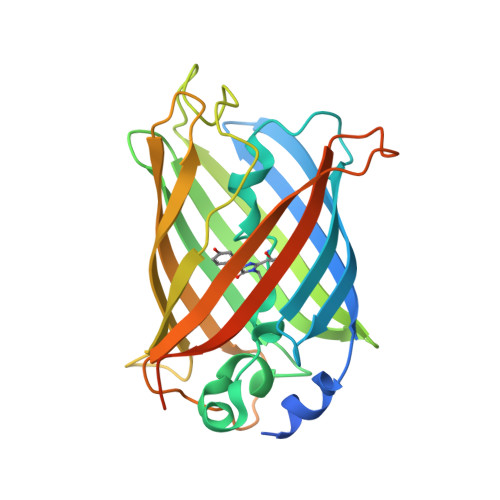
Figure 1 from AGGRESCAN3D (A3D): server for prediction of aggregation properties of protein structures | Semantic Scholar
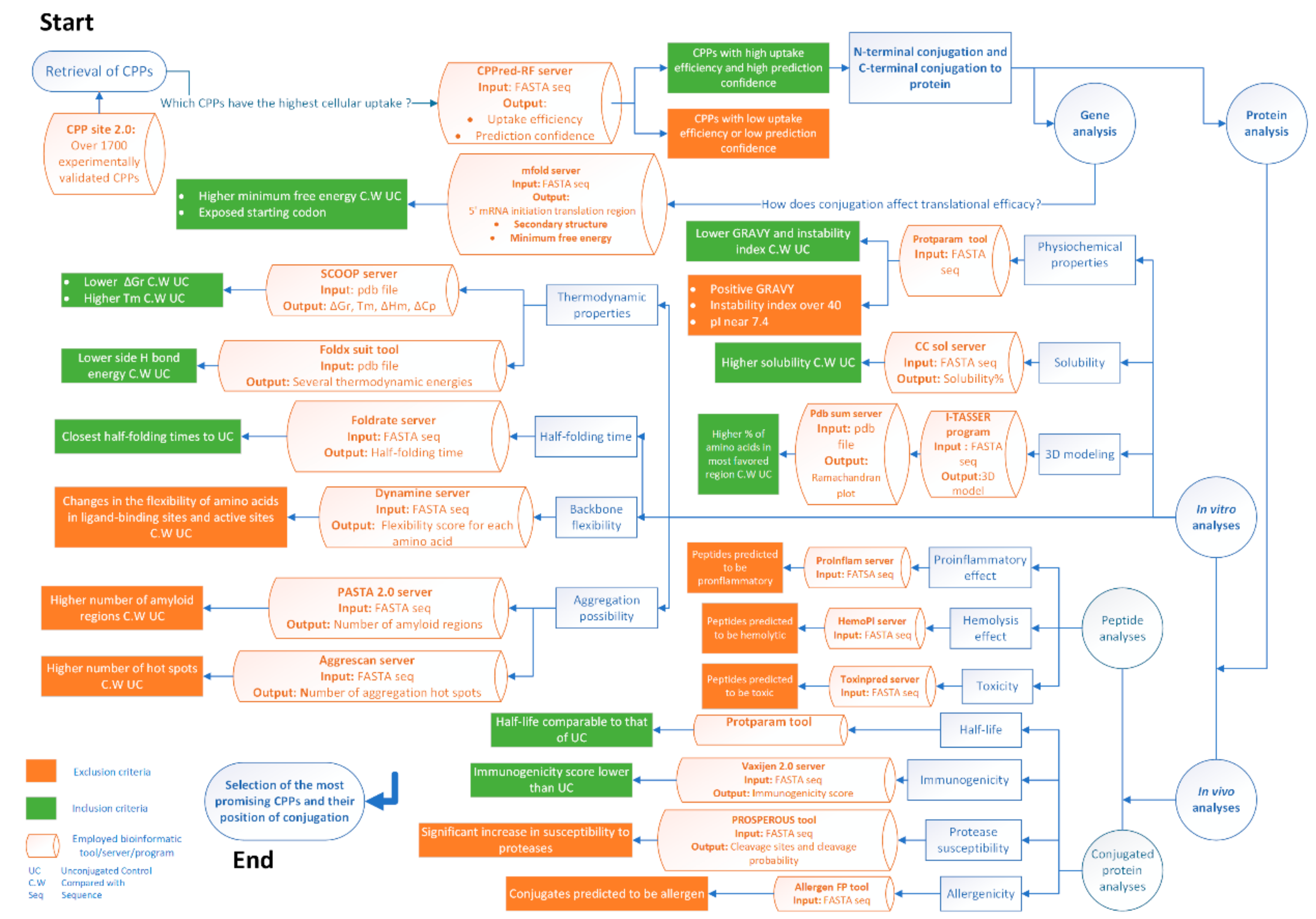
Molecules | Free Full-Text | Considerations on the Rational Design of Covalently Conjugated Cell-Penetrating Peptides (CPPs) for Intracellular Delivery of Proteins: A Guide to CPP Selection Using Glucarpidase as the Model Cargo

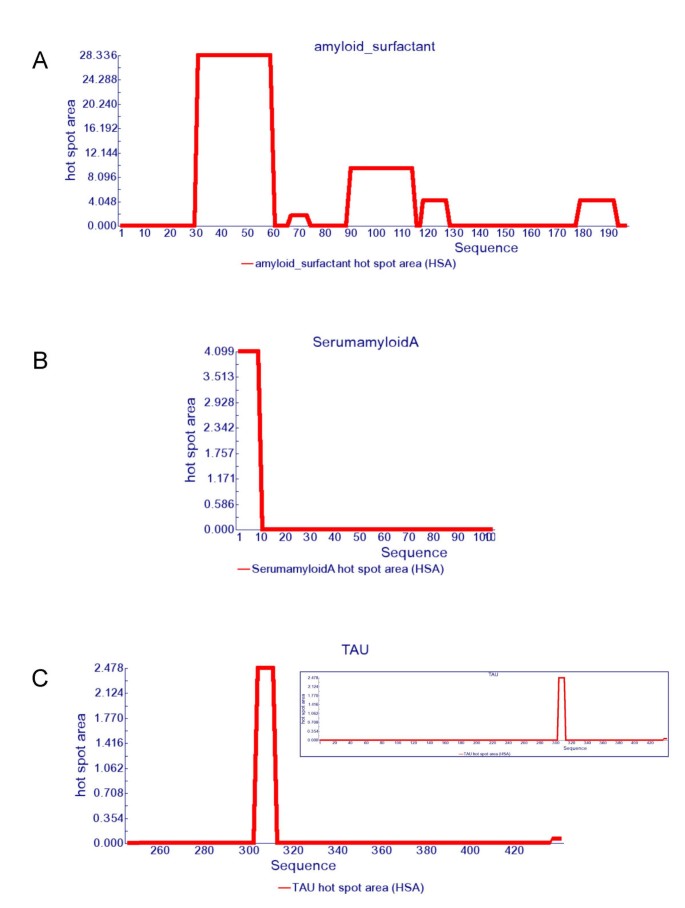
Aggregation-prone regions. (a) Aggregation-prone regions predicted by... | Download Scientific Diagram